methods
We put efforts on developing methodologies that can elucidate the functional mechanisms of high-molecular-weight proteins, establish structure-function relationship analyses that do not rely on mutants, and introduce intracellular structure-guided drug development. These various fundamental technologies will provide additional value to the structural and dynamical information revealed by NMR and promote its utilization.
Heteronuclear-direct detections

The application of solution NMR to high molecular weight (HMW) systems is still challenging, especially for those exceeding 100 kDa. The limitation mainly comes from signal losses due to fast transverse relaxation in HMW systems. While proton (1H) is the most commonly used as the detecting nucleus, the acceleration of relaxation in HMW systems is the most substantial among other nuclei.
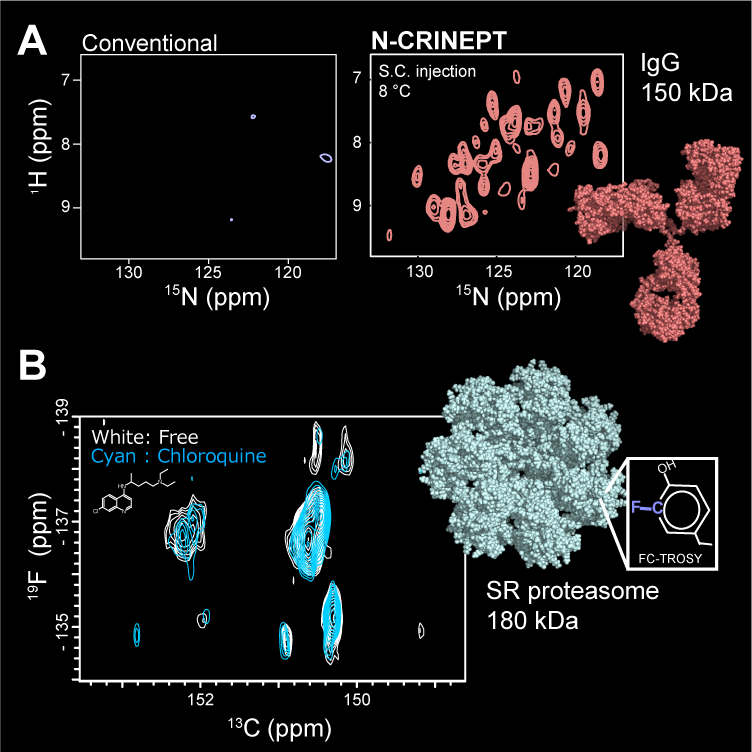
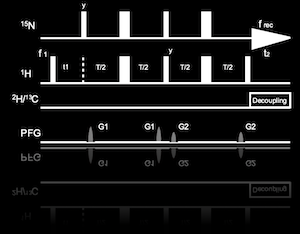
To overcome this limitation, we have developed heteronuclear-direct detection experiments that are applicable to the >100 kDa systems (Rec. Dev. Biomol. NMR, 2012; eMagRes, 2017). Those experiments use low γ-nuclei, such as 13C, and 15N for detection. Although the intrinsic sensitivities of these nuclei are lower than that of 1H, the slower relaxation yields a superior sensitivity in the non-deuterated or hard-to-deuterate conditions. Our latest examples are N-CRINPT (J. Med. Chem., 2020) and FC-TROSY (Nat. Methods., 2019). N-CRINEPT presents a structural fingerprint of an analog of a therapeutic monoclonal antibody at low storage temperature, under various prescription solution conditions (Panel A). FC-TROSY successfully provides the structural information from the aromatic sidechains of the 180-kDa α7 single-ring of the 20S proteasome core particle (Panel B). FC-TROSY is expected to provide a powerful background-free spectrum in in-cell NMR methods.
THz NMR

Manipulation of protein functions aids both biological research and disease treatments. However, the ways we have had for have been restricted to protein-targeting ligands, chemical modifications, and genetic manipulations. Application of electromagnetic waves at around terahertz (THz) frequency to proteins could be game-changing, as the THz radiation could modulate a protein function if it resonates with functionally relevant vibrational modes of the protein.
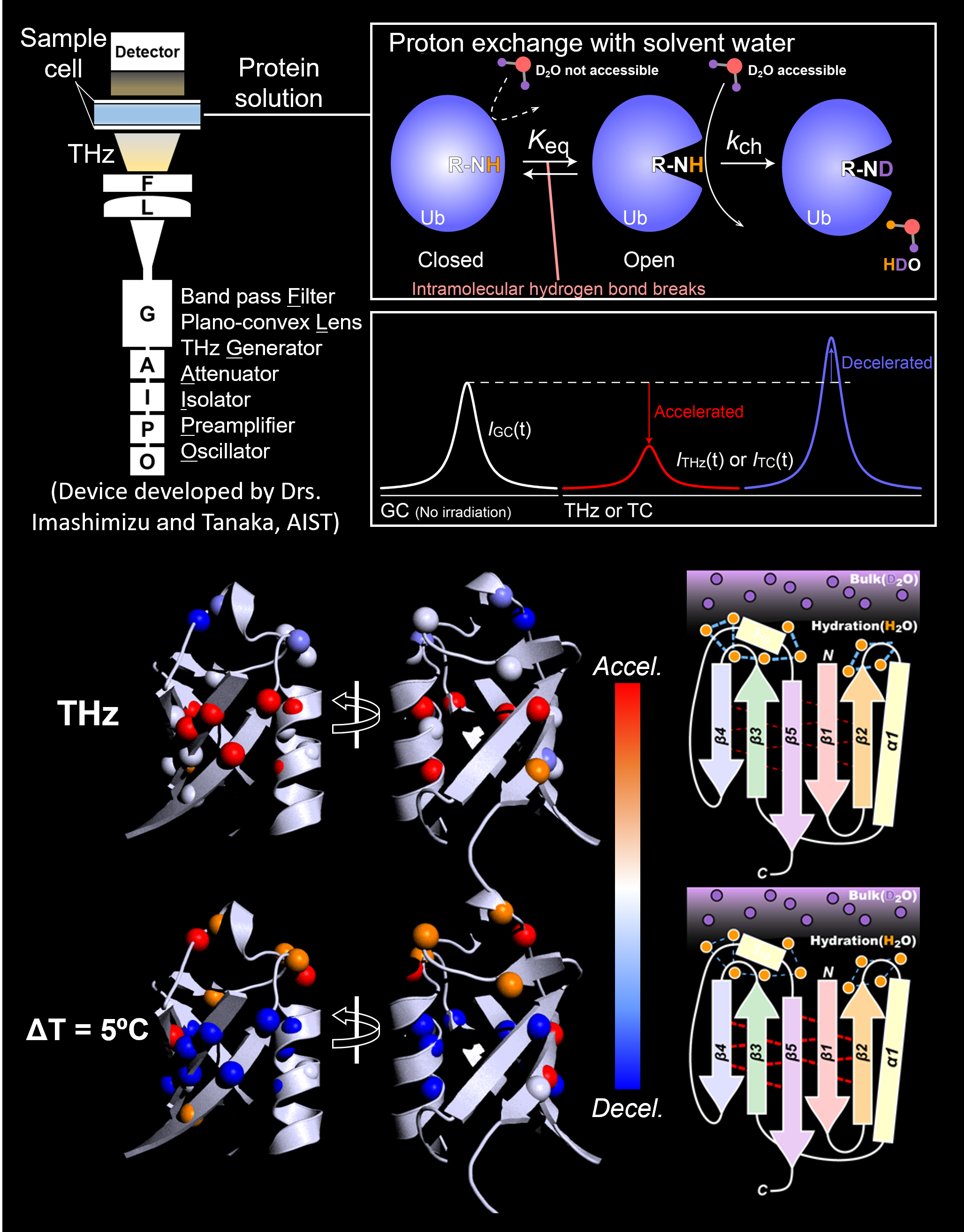
To make this practical, mechanistic insights into the effects of THz radiation on proteins would be prerequisite. We developed an experimental scheme to analyze the THz-induced changes in structure and dynamics of proteins in a residue-specific manner.
Transient, reversible effects of the THz radiation to the protein structures were encoded in NMR signal intensities via proton exchange with solvent water. With a strict control for excluding thermal effects, we successfully recapitulated the non-thermal effects of THz radiation on the hydration on a protein surface (Biophys J., 2021). We are now sophisticating the scheme to improve spatial temporal resolution, while examining the functional consequences of the THz-induced structural modulation.
Stable Isotope Labeling

In NMR analyses of proteins, the atoms of interest can be selectively observed by labeling appropriate atoms with stable isotopes such as 2H, 13C, and 15N. Especially, high-level deuteration of protons surrounding observed atoms enables sensitive detection of NMR signals in combination with the TROSY methods.
The isotope labeling strategy has been developed in E. coli expression systems, however, it is not that straight forwards in eukaryotic cells, such as insect cells and mammalian cells. Especially, since eukaryotic cells do not grow in heavy water (D2O), extensive deuteration are thought to be impossible.
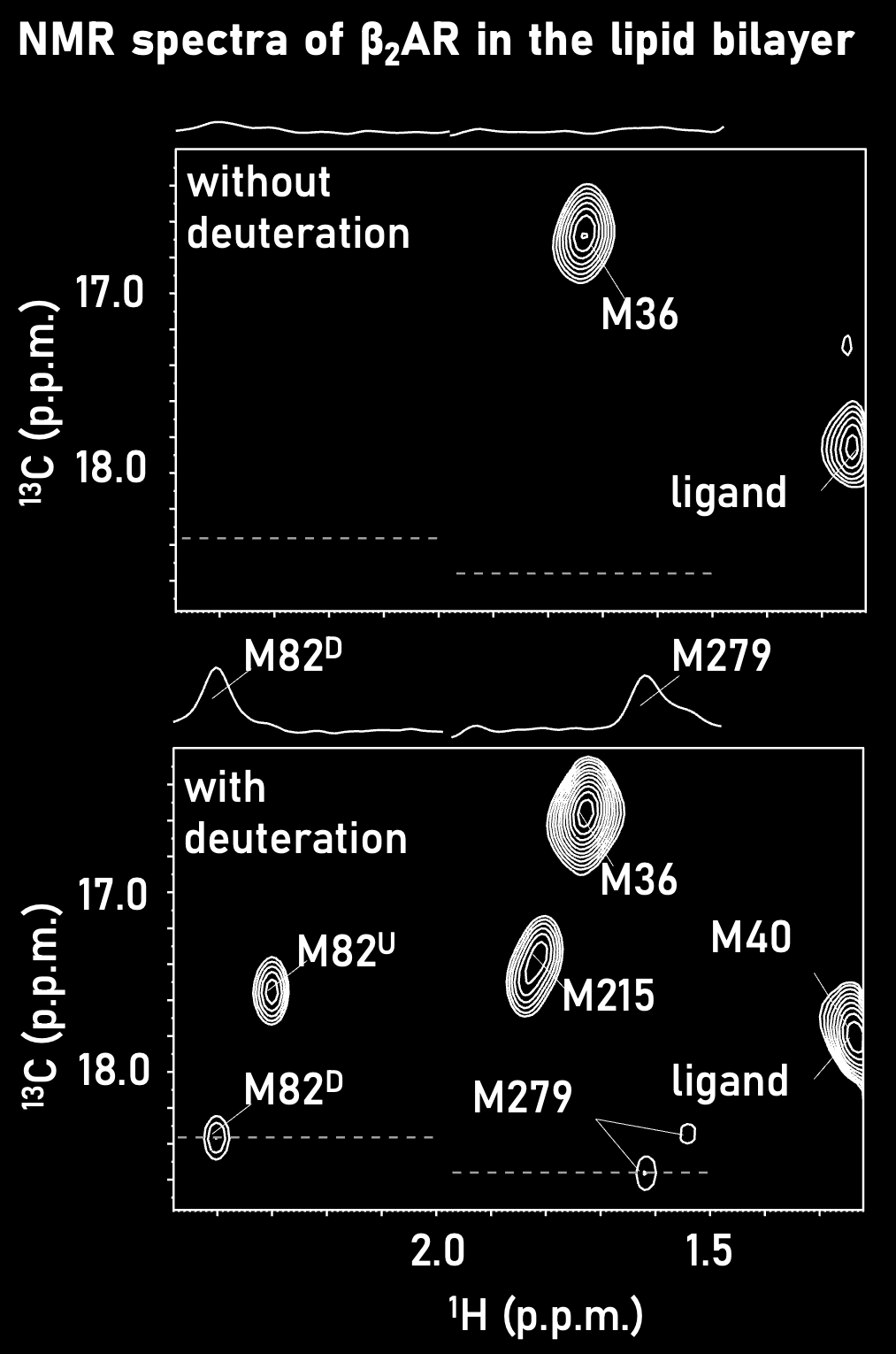
We have developed a strategy to selectively introduce methyl 13C-labeling with the high-level deuteration of surrounding amino acid residues in insect cells. Using this strategy, we have achieved a five-fold improvement in the sensitivity and succeeded the solution NMR analysis of GPCRs reconstituted in lipid bilayers for the first time (Angew. Chem. Int. Ed., 2014). The application of this technique has also led to the elucidation of the action mechanism of ligand-dependent ion channels in lipid bilayers (Proc. Natl. Acad. Sci. USA., 2016).By developing new stable isotope labeling techniques, we will push forward the NMR analysis of HMW proteins and membrane proteins, including important drug targets.
In cell drug discovery by NMR

In many cases, drug discovery starts from screening compounds that bind to the target in vitro. Followed by optimizing the structure and evaluating various factors required for the drug, a compound becomes a drug by spending ~ 10 years on average. However, it is not easy to integrate information obtained from different conditions. The critical difference between in vitro and in cell conditions sometimes leads to an unexpected difficulty, reducing the efficiency of the drug development process.
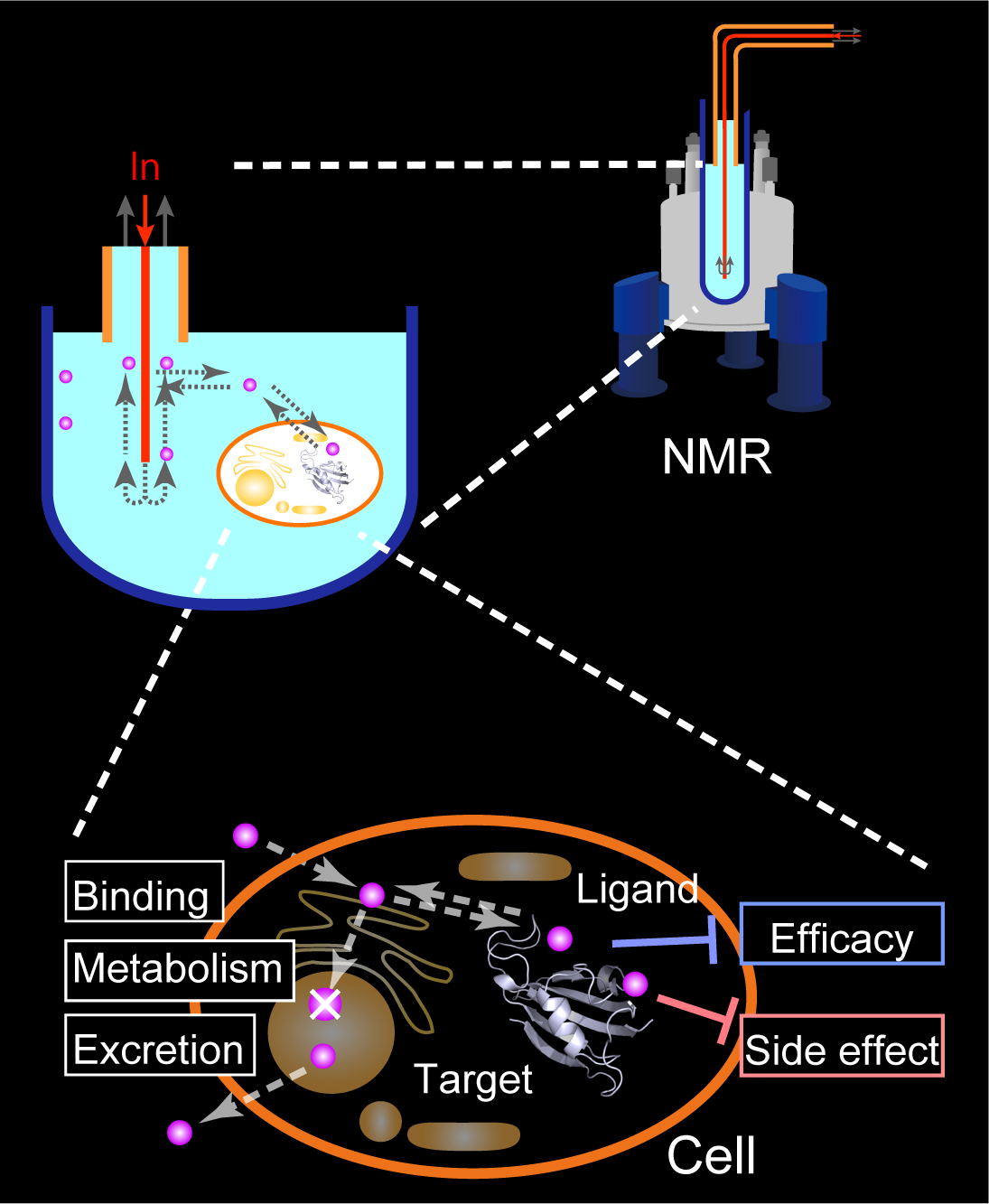
We think that carrying out the drug-discovery process in living cells using NMR would be beneficial. In the in-cell drug discovery (ICDD) by NMR, we will not only carry out the structure optimization of ligand but also analyze the cellular responses and toxicity under drug exposure using the living cells. The feature of NMR to conduct a wide range of analyses would benefit here. The development of the ICDD by NMR technology will greatly impact the early drug development process.